Session 4: Statistical Inference I - Parametric Tests
Event Studies in Finance and Economics - Summer School¶
Learning Objectives¶
By the end of this session, you will be able to:
Understand the assumptions underlying parametric event study tests
Implement and interpret the standard t-test for event studies
Apply the Patell (1976) standardized residual test
Use the Boehmer, Musumeci, and Poulsen (1991) test for event-induced variance
Distinguish between cross-sectional and time-series variance approaches
Evaluate the power and specification of different test statistics
1. Introduction: The Inference Problem¶
From Measurement to Inference¶
In Session 3, we learned how to measure abnormal returns. But measurement alone doesn’t tell us whether the event had a statistically significant effect. We need formal hypothesis tests.
The Null Hypothesis¶
Key Challenges¶
Estimation error: Parameters (, ) are estimated, not known
Cross-sectional correlation: Events may cluster in calendar time
Event-induced variance: The event itself may change volatility
Non-normality: Returns are often fat-tailed
Small samples: Limited number of events in many studies
This session focuses on parametric tests that assume (approximately) normal distributions. Session 5 will cover non-parametric alternatives.
2. Setup and Data¶
Source
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from scipy import stats
from scipy.special import comb
import yfinance as yf
import statsmodels.api as sm
from datetime import datetime, timedelta
from typing import List, Dict, Tuple, Optional
from dataclasses import dataclass
import warnings
warnings.filterwarnings('ignore')
plt.style.use('seaborn-v0_8-whitegrid')
sns.set_palette("husl")
print("Libraries loaded successfully!")Libraries loaded successfully!
Source
# We'll use a larger sample for better demonstration of test properties
EVENTS = [
# Tech earnings Q2-Q3 2023
{'ticker': 'AAPL', 'date': '2023-08-03', 'name': 'Apple Q3'},
{'ticker': 'MSFT', 'date': '2023-07-25', 'name': 'Microsoft Q4'},
{'ticker': 'GOOGL', 'date': '2023-07-25', 'name': 'Alphabet Q2'},
{'ticker': 'AMZN', 'date': '2023-08-03', 'name': 'Amazon Q2'},
{'ticker': 'META', 'date': '2023-07-26', 'name': 'Meta Q2'},
{'ticker': 'NVDA', 'date': '2023-08-23', 'name': 'Nvidia Q2'},
{'ticker': 'TSLA', 'date': '2023-07-19', 'name': 'Tesla Q2'},
{'ticker': 'AMD', 'date': '2023-08-01', 'name': 'AMD Q2'},
{'ticker': 'INTC', 'date': '2023-07-27', 'name': 'Intel Q2'},
{'ticker': 'CRM', 'date': '2023-08-30', 'name': 'Salesforce Q2'},
]
ESTIMATION_WINDOW = 120
GAP = 10
EVENT_WINDOW_PRE = 10
EVENT_WINDOW_POST = 10
print(f"Sample: {len(EVENTS)} tech earnings announcements")Sample: 10 tech earnings announcements
Source
@dataclass
class MarketModelResults:
"""Container for market model estimation results."""
alpha: float
beta: float
sigma: float
r_squared: float
n_obs: int
market_mean: float
market_var: float
residuals: np.ndarray
@dataclass
class EventStudyResult:
"""Container for single event study results."""
ticker: str
name: str
event_date: pd.Timestamp
model: MarketModelResults
event_data: pd.DataFrame
estimation_data: pd.DataFrame
def download_and_process_event(ticker: str, event_date: str, name: str,
est_window: int, gap: int,
pre: int, post: int) -> Optional[EventStudyResult]:
"""
Download data and estimate market model for a single event.
"""
try:
event_dt = pd.to_datetime(event_date)
start_date = event_dt - timedelta(days=int((est_window + gap + pre) * 1.5))
end_date = event_dt + timedelta(days=int(post * 2.5))
stock = yf.download(ticker, start=start_date, end=end_date, progress=False)['Close']
market = yf.download('^GSPC', start=start_date, end=end_date, progress=False)['Close']
df = pd.DataFrame({
'stock_price': stock.squeeze(),
'market_price': market.squeeze()
})
df['stock_ret'] = df['stock_price'].pct_change()
df['market_ret'] = df['market_price'].pct_change()
df = df.dropna()
if event_dt not in df.index:
idx = df.index.get_indexer([event_dt], method='nearest')[0]
event_dt = df.index[idx]
event_idx = df.index.get_loc(event_dt)
df['event_time'] = range(-event_idx, len(df) - event_idx)
# Split windows
est_end = -(gap + pre)
est_start = est_end - est_window
est_data = df[(df['event_time'] >= est_start) & (df['event_time'] < est_end)].copy()
evt_data = df[(df['event_time'] >= -pre) & (df['event_time'] <= post)].copy()
# Estimate market model
y = est_data['stock_ret'].values
x = est_data['market_ret'].values
X = sm.add_constant(x)
ols = sm.OLS(y, X).fit()
model = MarketModelResults(
alpha=ols.params[0],
beta=ols.params[1],
sigma=np.std(ols.resid, ddof=2),
r_squared=ols.rsquared,
n_obs=len(y),
market_mean=np.mean(x),
market_var=np.sum((x - np.mean(x))**2),
residuals=ols.resid
)
# Calculate abnormal returns
evt_data['expected_ret'] = model.alpha + model.beta * evt_data['market_ret']
evt_data['AR'] = evt_data['stock_ret'] - evt_data['expected_ret']
return EventStudyResult(
ticker=ticker,
name=name,
event_date=event_dt,
model=model,
event_data=evt_data,
estimation_data=est_data
)
except Exception as e:
print(f" {ticker}: FAILED - {e}")
return None
# Process all events
print("Processing events...")
event_results = []
for event in EVENTS:
result = download_and_process_event(
event['ticker'], event['date'], event['name'],
ESTIMATION_WINDOW, GAP, EVENT_WINDOW_PRE, EVENT_WINDOW_POST
)
if result:
event_results.append(result)
print(f" {result.ticker}: OK (beta={result.model.beta:.3f}, sigma={result.model.sigma:.4f})")
print(f"\nSuccessfully processed {len(event_results)} events")Processing events...
YF.download() has changed argument auto_adjust default to True
AAPL: OK (beta=1.127, sigma=0.0080)
MSFT: OK (beta=1.249, sigma=0.0140)
GOOGL: OK (beta=1.404, sigma=0.0166)
AMZN: OK (beta=1.521, sigma=0.0175)
META: OK (beta=1.891, sigma=0.0256)
NVDA: OK (beta=2.068, sigma=0.0305)
TSLA: OK (beta=2.163, sigma=0.0334)
AMD: OK (beta=1.770, sigma=0.0290)
INTC: OK (beta=1.304, sigma=0.0230)
CRM: OK (beta=1.078, sigma=0.0173)
Successfully processed 10 events
3. Test 1: Simple Cross-Sectional t-Test¶
The Simplest Approach¶
The most basic test treats individual CARs as independent draws and uses their cross-sectional distribution:
Where:
Assumptions¶
CARs are independent across firms
CARs are identically distributed
CARs are (approximately) normally distributed
Distribution¶
Under the null:
Source
def calculate_car(event_data: pd.DataFrame, tau1: int, tau2: int) -> float:
"""Calculate Cumulative Abnormal Return for a window."""
mask = (event_data['event_time'] >= tau1) & (event_data['event_time'] <= tau2)
return event_data.loc[mask, 'AR'].sum()
def cross_sectional_t_test(event_results: List[EventStudyResult],
tau1: int, tau2: int) -> Dict:
"""
Simple cross-sectional t-test for CAAR.
H0: E[CAR] = 0
Returns:
--------
Dictionary with test statistics
"""
# Calculate individual CARs
cars = np.array([calculate_car(r.event_data, tau1, tau2) for r in event_results])
N = len(cars)
# Sample statistics
caar = np.mean(cars)
s_car = np.std(cars, ddof=1)
se = s_car / np.sqrt(N)
# t-statistic
t_stat = caar / se if se > 0 else 0
# p-value (two-tailed)
p_value = 2 * (1 - stats.t.cdf(abs(t_stat), df=N-1))
return {
'test_name': 'Cross-Sectional t-Test',
'N': N,
'CAAR': caar,
'std_dev': s_car,
'std_error': se,
't_stat': t_stat,
'df': N - 1,
'p_value': p_value,
'individual_cars': cars
}
# Run test for standard windows
print("Cross-Sectional t-Test Results:")
print("="*70)
windows = [(-1, +1), (0, 0), (-5, +5), (-10, +10)]
cs_results = []
for tau1, tau2 in windows:
result = cross_sectional_t_test(event_results, tau1, tau2)
cs_results.append(result)
sig = '***' if result['p_value'] < 0.01 else '**' if result['p_value'] < 0.05 else '*' if result['p_value'] < 0.10 else ''
print(f"\nWindow [{tau1:+d}, {tau2:+d}]:")
print(f" CAAR = {result['CAAR']*100:+.3f}%")
print(f" Std. Dev. = {result['std_dev']*100:.3f}%")
print(f" t-stat = {result['t_stat']:.3f} {sig}")
print(f" p-value = {result['p_value']:.4f}")
print("\n" + "="*70)
print("Significance: * p<0.10, ** p<0.05, *** p<0.01")Cross-Sectional t-Test Results:
======================================================================
Window [-1, +1]:
CAAR = +0.959%
Std. Dev. = 6.380%
t-stat = 0.475
p-value = 0.6459
Window [+0, +0]:
CAAR = +0.600%
Std. Dev. = 1.223%
t-stat = 1.550
p-value = 0.1556
Window [-5, +5]:
CAAR = -0.761%
Std. Dev. = 6.559%
t-stat = -0.367
p-value = 0.7223
Window [-10, +10]:
CAAR = -5.226%
Std. Dev. = 7.930%
t-stat = -2.084 *
p-value = 0.0669
======================================================================
Significance: * p<0.10, ** p<0.05, *** p<0.01
Source
# Visualize individual CARs and distribution
fig, axes = plt.subplots(1, 2, figsize=(14, 5))
# Get CAR(-1,+1) results
result = cs_results[0] # (-1, +1) window
cars = result['individual_cars']
tickers = [r.ticker for r in event_results]
# Bar chart of individual CARs
ax1 = axes[0]
colors = ['green' if c >= 0 else 'red' for c in cars]
bars = ax1.bar(tickers, cars * 100, color=colors, alpha=0.7, edgecolor='black')
ax1.axhline(0, color='black', linewidth=0.5)
ax1.axhline(result['CAAR'] * 100, color='blue', linestyle='--',
label=f'CAAR = {result["CAAR"]*100:+.2f}%')
ax1.set_ylabel('CAR (-1, +1) %', fontsize=12)
ax1.set_title('Individual CARs', fontsize=14)
ax1.legend()
ax1.tick_params(axis='x', rotation=45)
# Distribution of CARs
ax2 = axes[1]
ax2.hist(cars * 100, bins=8, density=True, alpha=0.7, edgecolor='black', label='Empirical')
# Overlay normal distribution
x = np.linspace(cars.min()*100 - 2, cars.max()*100 + 2, 100)
y = stats.norm.pdf(x, result['CAAR']*100, result['std_dev']*100)
ax2.plot(x, y, 'r-', linewidth=2, label='Normal fit')
ax2.axvline(0, color='black', linestyle='-', linewidth=0.5)
ax2.axvline(result['CAAR']*100, color='blue', linestyle='--', label='Mean')
ax2.set_xlabel('CAR (%)', fontsize=12)
ax2.set_ylabel('Density', fontsize=12)
ax2.set_title('Distribution of CARs', fontsize=14)
ax2.legend()
plt.tight_layout()
plt.show()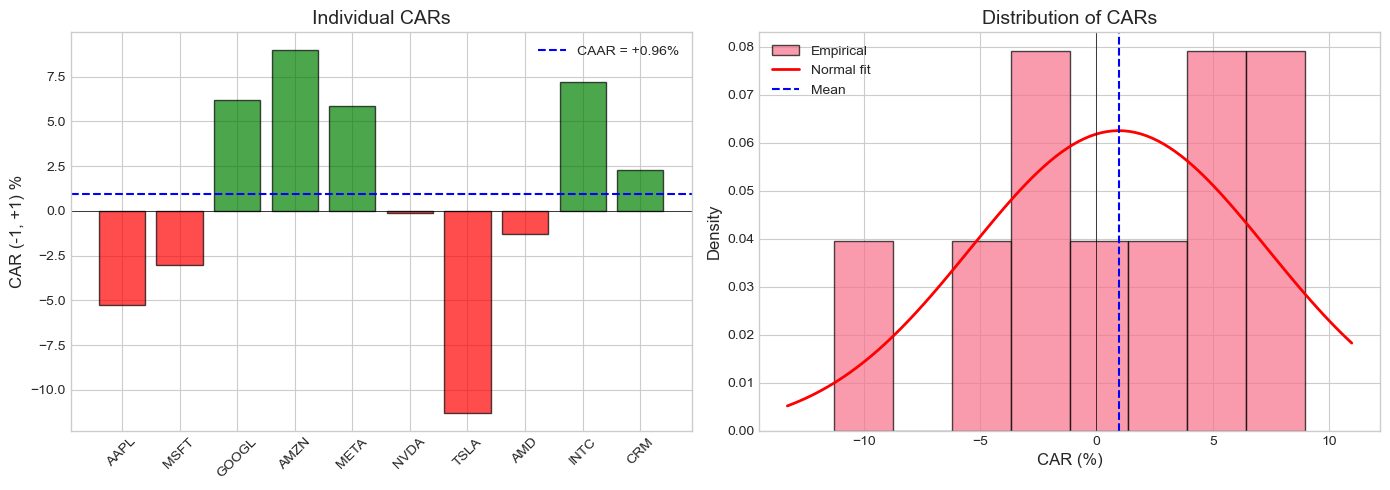
4. Test 2: Time-Series Standard Error Approach¶
Alternative Variance Estimation¶
Instead of using cross-sectional variation, we can use the time-series variance from the estimation period:
Where is the length of the event window.
Aggregated Test Statistic¶
Comparison¶
| Approach | Variance Source | Pros | Cons |
|---|---|---|---|
| Cross-sectional | Event window | Captures heterogeneity | Needs many events |
| Time-series | Estimation window | Works with few events | Assumes stable variance |
Source
def calculate_car_variance_ts(model: MarketModelResults,
event_data: pd.DataFrame,
tau1: int, tau2: int) -> float:
"""
Calculate time-series variance of CAR.
Includes adjustment for estimation error in alpha and beta.
"""
mask = (event_data['event_time'] >= tau1) & (event_data['event_time'] <= tau2)
window_data = event_data[mask]
L = len(window_data)
T = model.n_obs
sigma_sq = model.sigma ** 2
Rm_bar = model.market_mean
sum_sq = model.market_var
# Sum of market return deviations in event window
Rm_event = window_data['market_ret'].values
sum_dev_sq = np.sum((Rm_event - Rm_bar) ** 2)
# Variance formula (Campbell, Lo, MacKinlay 1997, eq. 4.8)
var = sigma_sq * (L + L**2/T + sum_dev_sq/sum_sq)
return var
def time_series_t_test(event_results: List[EventStudyResult],
tau1: int, tau2: int) -> Dict:
"""
Time-series variance t-test for CAAR.
Uses estimation period variance instead of cross-sectional variance.
"""
cars = []
car_vars = []
for r in event_results:
car = calculate_car(r.event_data, tau1, tau2)
var = calculate_car_variance_ts(r.model, r.event_data, tau1, tau2)
cars.append(car)
car_vars.append(var)
cars = np.array(cars)
car_vars = np.array(car_vars)
N = len(cars)
caar = np.mean(cars)
# Time-series aggregated variance
# Assuming independence: Var(CAAR) = (1/N^2) * sum(Var(CAR_i))
var_caar = np.sum(car_vars) / (N ** 2)
se_caar = np.sqrt(var_caar)
t_stat = caar / se_caar if se_caar > 0 else 0
# Approximate df (conservative)
df = N * (ESTIMATION_WINDOW - 2)
p_value = 2 * (1 - stats.t.cdf(abs(t_stat), df=df))
return {
'test_name': 'Time-Series t-Test',
'N': N,
'CAAR': caar,
'var_caar': var_caar,
'std_error': se_caar,
't_stat': t_stat,
'df': df,
'p_value': p_value,
'individual_cars': cars,
'individual_vars': car_vars
}
# Compare cross-sectional and time-series tests
print("Cross-Sectional vs Time-Series t-Tests:")
print("="*80)
comparison_data = []
for tau1, tau2 in windows:
cs = cross_sectional_t_test(event_results, tau1, tau2)
ts = time_series_t_test(event_results, tau1, tau2)
comparison_data.append({
'Window': f'[{tau1:+d},{tau2:+d}]',
'CAAR': cs['CAAR'],
'CS_SE': cs['std_error'],
'CS_t': cs['t_stat'],
'CS_p': cs['p_value'],
'TS_SE': ts['std_error'],
'TS_t': ts['t_stat'],
'TS_p': ts['p_value']
})
comp_df = pd.DataFrame(comparison_data)
display_comp = comp_df.copy()
display_comp['CAAR'] = display_comp['CAAR'].apply(lambda x: f"{x*100:+.3f}%")
display_comp['CS_SE'] = display_comp['CS_SE'].apply(lambda x: f"{x*100:.3f}%")
display_comp['TS_SE'] = display_comp['TS_SE'].apply(lambda x: f"{x*100:.3f}%")
for col in ['CS_t', 'TS_t']:
display_comp[col] = display_comp[col].apply(lambda x: f"{x:+.3f}")
for col in ['CS_p', 'TS_p']:
display_comp[col] = display_comp[col].apply(lambda x: f"{x:.4f}")
print(display_comp.to_string(index=False))
print("\nCS = Cross-Sectional, TS = Time-Series")Cross-Sectional vs Time-Series t-Tests:
================================================================================
Window CAAR CS_SE CS_t CS_p TS_SE TS_t TS_p
[-1,+1] +0.959% 2.018% +0.475 0.6459 1.271% +0.755 0.4506
[+0,+0] +0.600% 0.387% +1.550 0.1556 0.727% +0.825 0.4095
[-5,+5] -0.761% 2.074% -0.367 0.7223 2.509% -0.303 0.7618
[-10,+10] -5.226% 2.508% -2.084 0.0669 3.595% -1.454 0.1463
CS = Cross-Sectional, TS = Time-Series
5. Test 3: Patell (1976) Standardized Residual Test¶
Motivation¶
The simple tests treat all firms equally, but firms have different estimation period variances. Patell’s test standardizes each firm’s abnormal return by its own standard error.
Standardized Abnormal Return¶
Where:
Standardized CAR¶
Where:
Patell Test Statistic¶
Under the null:
Source
def calculate_sar(event_data: pd.DataFrame, model: MarketModelResults) -> np.ndarray:
"""
Calculate Standardized Abnormal Returns.
"""
T = model.n_obs
sigma = model.sigma
Rm_bar = model.market_mean
sum_sq = model.market_var
Rm = event_data['market_ret'].values
AR = event_data['AR'].values
# Standard error for each AR
S_AR = sigma * np.sqrt(1 + 1/T + (Rm - Rm_bar)**2 / sum_sq)
# Standardized AR
SAR = AR / S_AR
return SAR, S_AR
def calculate_scar(event_data: pd.DataFrame, model: MarketModelResults,
tau1: int, tau2: int) -> Tuple[float, float]:
"""
Calculate Standardized Cumulative Abnormal Return.
Returns: (SCAR, CAR)
"""
mask = (event_data['event_time'] >= tau1) & (event_data['event_time'] <= tau2)
window_data = event_data[mask].copy()
_, S_AR = calculate_sar(window_data, model)
CAR = window_data['AR'].sum()
S_CAR = np.sqrt(np.sum(S_AR**2))
SCAR = CAR / S_CAR if S_CAR > 0 else 0
return SCAR, CAR
def patell_test(event_results: List[EventStudyResult],
tau1: int, tau2: int) -> Dict:
"""
Patell (1976) Standardized Residual Test.
Aggregates standardized CARs across firms.
"""
scars = []
cars = []
adjustment_factors = []
for r in event_results:
scar, car = calculate_scar(r.event_data, r.model, tau1, tau2)
T = r.model.n_obs
# Patell adjustment factor
adj = np.sqrt((T - 4) / (T - 2))
scars.append(scar)
cars.append(car)
adjustment_factors.append(adj)
scars = np.array(scars)
cars = np.array(cars)
adj_factors = np.array(adjustment_factors)
N = len(scars)
# Patell Z-statistic
# Z = (1/sqrt(N)) * sum(SCAR_i * adjustment_i)
Z_patell = np.sum(scars * adj_factors) / np.sqrt(N)
# p-value (standard normal)
p_value = 2 * (1 - stats.norm.cdf(abs(Z_patell)))
return {
'test_name': 'Patell (1976)',
'N': N,
'CAAR': np.mean(cars),
'mean_SCAR': np.mean(scars),
'Z_stat': Z_patell,
'p_value': p_value,
'individual_scars': scars,
'individual_cars': cars
}
# Run Patell test
print("Patell (1976) Standardized Residual Test:")
print("="*70)
patell_results = []
for tau1, tau2 in windows:
result = patell_test(event_results, tau1, tau2)
patell_results.append(result)
sig = '***' if result['p_value'] < 0.01 else '**' if result['p_value'] < 0.05 else '*' if result['p_value'] < 0.10 else ''
print(f"\nWindow [{tau1:+d}, {tau2:+d}]:")
print(f" CAAR = {result['CAAR']*100:+.3f}%")
print(f" Mean SCAR = {result['mean_SCAR']:+.3f}")
print(f" Z-stat = {result['Z_stat']:+.3f} {sig}")
print(f" p-value = {result['p_value']:.4f}")Patell (1976) Standardized Residual Test:
======================================================================
Window [-1, +1]:
CAAR = +0.959%
Mean SCAR = +0.175
Z-stat = +0.550
p-value = 0.5825
Window [+0, +0]:
CAAR = +0.600%
Mean SCAR = +0.252
Z-stat = +0.790
p-value = 0.4298
Window [-5, +5]:
CAAR = -0.761%
Mean SCAR = -0.244
Z-stat = -0.764
p-value = 0.4449
Window [-10, +10]:
CAAR = -5.226%
Mean SCAR = -0.592
Z-stat = -1.858 *
p-value = 0.0632
6. Test 4: Boehmer, Musumeci, and Poulsen (1991) Test¶
The Event-Induced Variance Problem¶
The Patell test can be severely misspecified when events induce changes in variance. If the event increases volatility, Patell’s test will over-reject the null.
BMP Solution¶
Use the cross-sectional variance of standardized abnormal returns instead of assuming unit variance:
Where:
Why It Works¶
If event-induced variance affects all firms similarly, the cross-sectional variance of SCARs will capture it, providing a more robust test.
Source
def bmp_test(event_results: List[EventStudyResult],
tau1: int, tau2: int) -> Dict:
"""
Boehmer, Musumeci, and Poulsen (1991) Test.
Uses cross-sectional standard error of SCARs to account for
event-induced variance changes.
"""
scars = []
cars = []
for r in event_results:
scar, car = calculate_scar(r.event_data, r.model, tau1, tau2)
scars.append(scar)
cars.append(car)
scars = np.array(scars)
cars = np.array(cars)
N = len(scars)
# BMP statistic uses cross-sectional variance of SCARs
mean_scar = np.mean(scars)
std_scar = np.std(scars, ddof=1)
se_scar = std_scar / np.sqrt(N)
# Z-statistic
Z_bmp = mean_scar / se_scar if se_scar > 0 else 0
# Under null with event-induced variance, this should be t-distributed
p_value = 2 * (1 - stats.t.cdf(abs(Z_bmp), df=N-1))
return {
'test_name': 'BMP (1991)',
'N': N,
'CAAR': np.mean(cars),
'mean_SCAR': mean_scar,
'std_SCAR': std_scar,
'se_SCAR': se_scar,
't_stat': Z_bmp,
'df': N - 1,
'p_value': p_value,
'individual_scars': scars,
'individual_cars': cars
}
# Run BMP test
print("Boehmer-Musumeci-Poulsen (1991) Test:")
print("="*70)
bmp_results = []
for tau1, tau2 in windows:
result = bmp_test(event_results, tau1, tau2)
bmp_results.append(result)
sig = '***' if result['p_value'] < 0.01 else '**' if result['p_value'] < 0.05 else '*' if result['p_value'] < 0.10 else ''
print(f"\nWindow [{tau1:+d}, {tau2:+d}]:")
print(f" CAAR = {result['CAAR']*100:+.3f}%")
print(f" Mean SCAR = {result['mean_SCAR']:+.3f}")
print(f" Std. SCAR = {result['std_SCAR']:.3f}")
print(f" t-stat = {result['t_stat']:+.3f} {sig}")
print(f" p-value = {result['p_value']:.4f}")Boehmer-Musumeci-Poulsen (1991) Test:
======================================================================
Window [-1, +1]:
CAAR = +0.959%
Mean SCAR = +0.175
Std. SCAR = 2.048
t-stat = +0.271
p-value = 0.7927
Window [+0, +0]:
CAAR = +0.600%
Mean SCAR = +0.252
Std. SCAR = 0.565
t-stat = +1.410
p-value = 0.1923
Window [-5, +5]:
CAAR = -0.761%
Mean SCAR = -0.244
Std. SCAR = 1.335
t-stat = -0.577
p-value = 0.5781
Window [-10, +10]:
CAAR = -5.226%
Mean SCAR = -0.592
Std. SCAR = 1.044
t-stat = -1.794
p-value = 0.1064
Source
# Compare Patell vs BMP
print("\nPatell vs BMP Comparison:")
print("="*80)
print("\nIf Std. SCAR > 1, events are increasing variance (BMP is more conservative)")
print("If Std. SCAR < 1, events are decreasing variance (BMP is more powerful)\n")
pvb_data = []
for i, (tau1, tau2) in enumerate(windows):
patell = patell_results[i]
bmp = bmp_results[i]
pvb_data.append({
'Window': f'[{tau1:+d},{tau2:+d}]',
'Patell_Z': patell['Z_stat'],
'Patell_p': patell['p_value'],
'BMP_t': bmp['t_stat'],
'BMP_p': bmp['p_value'],
'Std_SCAR': bmp['std_SCAR'],
'Variance_Effect': 'Increase' if bmp['std_SCAR'] > 1.1 else 'Decrease' if bmp['std_SCAR'] < 0.9 else 'Neutral'
})
pvb_df = pd.DataFrame(pvb_data)
display_pvb = pvb_df.copy()
for col in ['Patell_Z', 'BMP_t']:
display_pvb[col] = display_pvb[col].apply(lambda x: f"{x:+.3f}")
for col in ['Patell_p', 'BMP_p']:
display_pvb[col] = display_pvb[col].apply(lambda x: f"{x:.4f}")
display_pvb['Std_SCAR'] = display_pvb['Std_SCAR'].apply(lambda x: f"{x:.3f}")
print(display_pvb.to_string(index=False))
Patell vs BMP Comparison:
================================================================================
If Std. SCAR > 1, events are increasing variance (BMP is more conservative)
If Std. SCAR < 1, events are decreasing variance (BMP is more powerful)
Window Patell_Z Patell_p BMP_t BMP_p Std_SCAR Variance_Effect
[-1,+1] +0.550 0.5825 +0.271 0.7927 2.048 Increase
[+0,+0] +0.790 0.4298 +1.410 0.1923 0.565 Decrease
[-5,+5] -0.764 0.4449 -0.577 0.5781 1.335 Increase
[-10,+10] -1.858 0.0632 -1.794 0.1064 1.044 Neutral
7. Test 5: Kolari and Pynnönen (2010) Adjusted BMP¶
Cross-Correlation Problem¶
When events cluster in calendar time, abnormal returns may be cross-sectionally correlated. This violates the independence assumption and inflates test statistics.
Adjustment¶
Kolari and Pynnönen (2010) propose adjusting the BMP test for cross-correlation:
Where is the average cross-correlation of abnormal returns in the estimation period.
Source
def estimate_cross_correlation(event_results: List[EventStudyResult]) -> float:
"""
Estimate average cross-correlation of residuals.
Uses estimation period residuals to estimate cross-sectional correlation.
"""
# Collect residuals (may have different lengths/dates, so we use pairwise correlation)
residuals_list = [r.model.residuals for r in event_results]
N = len(residuals_list)
if N < 2:
return 0.0
# Calculate pairwise correlations
correlations = []
for i in range(N):
for j in range(i+1, N):
# Use minimum common length
min_len = min(len(residuals_list[i]), len(residuals_list[j]))
if min_len > 10:
corr = np.corrcoef(residuals_list[i][:min_len],
residuals_list[j][:min_len])[0, 1]
if not np.isnan(corr):
correlations.append(corr)
return np.mean(correlations) if correlations else 0.0
def kolari_pynnonen_test(event_results: List[EventStudyResult],
tau1: int, tau2: int) -> Dict:
"""
Kolari and Pynnönen (2010) adjusted BMP test.
Adjusts for cross-sectional correlation of abnormal returns.
"""
# First get BMP results
bmp = bmp_test(event_results, tau1, tau2)
N = bmp['N']
t_bmp = bmp['t_stat']
# Estimate cross-correlation
r_bar = estimate_cross_correlation(event_results)
# Adjustment factor
if r_bar >= 1/(1-N): # Avoid negative under radical
adj_factor = 0.01 # Very small adjustment
else:
adj_factor = np.sqrt((1 - r_bar) / (1 + (N-1) * r_bar))
# Adjusted t-statistic
t_adj = t_bmp * adj_factor
# p-value
p_value = 2 * (1 - stats.t.cdf(abs(t_adj), df=N-1))
return {
'test_name': 'Kolari-Pynnönen (2010)',
'N': N,
'CAAR': bmp['CAAR'],
'mean_SCAR': bmp['mean_SCAR'],
't_BMP': t_bmp,
'r_bar': r_bar,
'adj_factor': adj_factor,
't_adjusted': t_adj,
'p_value': p_value
}
# Run Kolari-Pynnönen test
print("Kolari-Pynnönen (2010) Adjusted Test:")
print("="*70)
# First show cross-correlation estimate
r_bar = estimate_cross_correlation(event_results)
print(f"\nEstimated average cross-correlation: {r_bar:.4f}")
print("(High correlation requires larger adjustment)\n")
kp_results = []
for tau1, tau2 in windows:
result = kolari_pynnonen_test(event_results, tau1, tau2)
kp_results.append(result)
sig = '***' if result['p_value'] < 0.01 else '**' if result['p_value'] < 0.05 else '*' if result['p_value'] < 0.10 else ''
print(f"Window [{tau1:+d}, {tau2:+d}]:")
print(f" BMP t-stat = {result['t_BMP']:+.3f}")
print(f" Adjustment factor = {result['adj_factor']:.3f}")
print(f" Adjusted t-stat = {result['t_adjusted']:+.3f} {sig}")
print(f" p-value = {result['p_value']:.4f}\n")Kolari-Pynnönen (2010) Adjusted Test:
======================================================================
Estimated average cross-correlation: 0.0203
(High correlation requires larger adjustment)
Window [-1, +1]:
BMP t-stat = +0.271
Adjustment factor = 0.010
Adjusted t-stat = +0.003
p-value = 0.9979
Window [+0, +0]:
BMP t-stat = +1.410
Adjustment factor = 0.010
Adjusted t-stat = +0.014
p-value = 0.9891
Window [-5, +5]:
BMP t-stat = -0.577
Adjustment factor = 0.010
Adjusted t-stat = -0.006
p-value = 0.9955
Window [-10, +10]:
BMP t-stat = -1.794
Adjustment factor = 0.010
Adjusted t-stat = -0.018
p-value = 0.9861
8. Comprehensive Test Comparison¶
Let’s compare all parametric tests side by side.
Source
def run_all_parametric_tests(event_results: List[EventStudyResult],
tau1: int, tau2: int) -> pd.DataFrame:
"""
Run all parametric tests and return comparison table.
"""
cs = cross_sectional_t_test(event_results, tau1, tau2)
ts = time_series_t_test(event_results, tau1, tau2)
patell = patell_test(event_results, tau1, tau2)
bmp = bmp_test(event_results, tau1, tau2)
kp = kolari_pynnonen_test(event_results, tau1, tau2)
results = [
{'Test': 'Cross-Sectional t', 'Statistic': cs['t_stat'], 'p-value': cs['p_value']},
{'Test': 'Time-Series t', 'Statistic': ts['t_stat'], 'p-value': ts['p_value']},
{'Test': 'Patell Z', 'Statistic': patell['Z_stat'], 'p-value': patell['p_value']},
{'Test': 'BMP t', 'Statistic': bmp['t_stat'], 'p-value': bmp['p_value']},
{'Test': 'Kolari-Pynnönen t', 'Statistic': kp['t_adjusted'], 'p-value': kp['p_value']},
]
return pd.DataFrame(results)
# Create comprehensive comparison
print("\n" + "="*90)
print("COMPREHENSIVE PARAMETRIC TEST COMPARISON")
print("="*90)
for tau1, tau2 in windows:
print(f"\n\nWindow [{tau1:+d}, {tau2:+d}]")
print("-" * 60)
# Get CAAR for reference
cars = [calculate_car(r.event_data, tau1, tau2) for r in event_results]
caar = np.mean(cars)
print(f"CAAR = {caar*100:+.3f}% (N = {len(event_results)})\n")
results = run_all_parametric_tests(event_results, tau1, tau2)
# Add significance markers
def sig_marker(p):
if p < 0.01: return '***'
elif p < 0.05: return '**'
elif p < 0.10: return '*'
else: return ''
results['Sig'] = results['p-value'].apply(sig_marker)
results['Statistic'] = results['Statistic'].apply(lambda x: f"{x:+.3f}")
results['p-value'] = results['p-value'].apply(lambda x: f"{x:.4f}")
print(results.to_string(index=False))
==========================================================================================
COMPREHENSIVE PARAMETRIC TEST COMPARISON
==========================================================================================
Window [-1, +1]
------------------------------------------------------------
CAAR = +0.959% (N = 10)
Test Statistic p-value Sig
Cross-Sectional t +0.475 0.6459
Time-Series t +0.755 0.4506
Patell Z +0.550 0.5825
BMP t +0.271 0.7927
Kolari-Pynnönen t +0.003 0.9979
Window [+0, +0]
------------------------------------------------------------
CAAR = +0.600% (N = 10)
Test Statistic p-value Sig
Cross-Sectional t +1.550 0.1556
Time-Series t +0.825 0.4095
Patell Z +0.790 0.4298
BMP t +1.410 0.1923
Kolari-Pynnönen t +0.014 0.9891
Window [-5, +5]
------------------------------------------------------------
CAAR = -0.761% (N = 10)
Test Statistic p-value Sig
Cross-Sectional t -0.367 0.7223
Time-Series t -0.303 0.7618
Patell Z -0.764 0.4449
BMP t -0.577 0.5781
Kolari-Pynnönen t -0.006 0.9955
Window [-10, +10]
------------------------------------------------------------
CAAR = -5.226% (N = 10)
Test Statistic p-value Sig
Cross-Sectional t -2.084 0.0669 *
Time-Series t -1.454 0.1463
Patell Z -1.858 0.0632 *
BMP t -1.794 0.1064
Kolari-Pynnönen t -0.018 0.9861
Source
# Visualize test statistics comparison
fig, axes = plt.subplots(1, 2, figsize=(14, 5))
# Collect test statistics for all windows
test_names = ['CS t', 'TS t', 'Patell Z', 'BMP t', 'KP t']
window_labels = [f'[{t1:+d},{t2:+d}]' for t1, t2 in windows]
stats_matrix = np.zeros((len(windows), len(test_names)))
for i, (tau1, tau2) in enumerate(windows):
cs = cross_sectional_t_test(event_results, tau1, tau2)
ts = time_series_t_test(event_results, tau1, tau2)
patell = patell_test(event_results, tau1, tau2)
bmp = bmp_test(event_results, tau1, tau2)
kp = kolari_pynnonen_test(event_results, tau1, tau2)
stats_matrix[i, :] = [cs['t_stat'], ts['t_stat'], patell['Z_stat'],
bmp['t_stat'], kp['t_adjusted']]
# Heatmap of test statistics
ax1 = axes[0]
im = ax1.imshow(stats_matrix.T, cmap='RdYlGn', aspect='auto', vmin=-3, vmax=3)
ax1.set_xticks(range(len(windows)))
ax1.set_xticklabels(window_labels)
ax1.set_yticks(range(len(test_names)))
ax1.set_yticklabels(test_names)
ax1.set_xlabel('Event Window')
ax1.set_title('Test Statistics Across Windows')
# Add value annotations
for i in range(len(test_names)):
for j in range(len(windows)):
ax1.text(j, i, f'{stats_matrix[j, i]:.2f}', ha='center', va='center', fontsize=9)
plt.colorbar(im, ax=ax1, label='Test Statistic')
# Critical value lines for reference
ax2 = axes[1]
x = np.arange(len(windows))
width = 0.15
for i, test in enumerate(test_names):
ax2.bar(x + i*width, stats_matrix[:, i], width, label=test, alpha=0.8)
ax2.axhline(1.96, color='red', linestyle='--', alpha=0.7, label='5% critical')
ax2.axhline(-1.96, color='red', linestyle='--', alpha=0.7)
ax2.axhline(1.645, color='orange', linestyle=':', alpha=0.7, label='10% critical')
ax2.axhline(-1.645, color='orange', linestyle=':', alpha=0.7)
ax2.set_xticks(x + 2*width)
ax2.set_xticklabels(window_labels)
ax2.set_xlabel('Event Window')
ax2.set_ylabel('Test Statistic')
ax2.set_title('Test Statistics Comparison')
ax2.legend(bbox_to_anchor=(1.02, 1), loc='upper left', fontsize=8)
plt.tight_layout()
plt.show()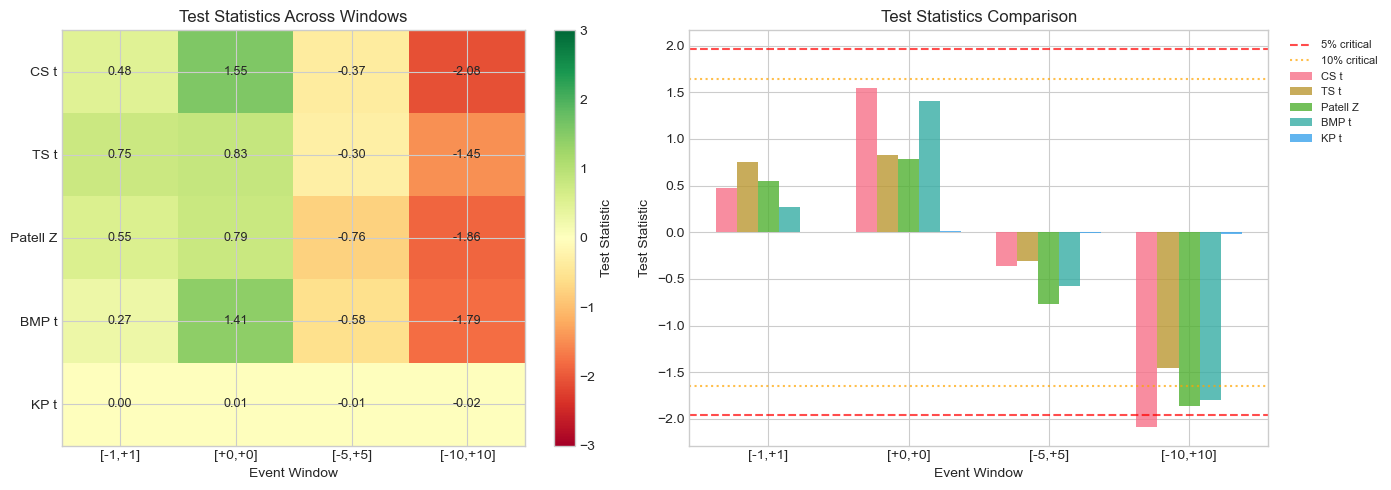
9. Power Analysis: Monte Carlo Simulation¶
Understanding Test Power¶
Power = Probability of rejecting when it is false
We want tests with:
Correct size (reject at stated significance level under )
High power (detect true effects)
Simulation Setup¶
Generate synthetic abnormal returns with known properties
Apply different tests
Measure rejection rates under (size) and (power)
Source
def simulate_event_study(N: int, T_est: int, L_event: int,
true_car: float = 0.0,
sigma: float = 0.02,
event_variance_multiplier: float = 1.0) -> List[Dict]:
"""
Simulate event study data for power analysis.
Parameters:
-----------
N : int
Number of events
T_est : int
Estimation window length
L_event : int
Event window length
true_car : float
True cumulative abnormal return (0 under null)
sigma : float
Residual standard deviation
event_variance_multiplier : float
Multiplier for event window variance (>1 means event-induced variance)
Returns:
--------
List of simulated event results
"""
results = []
# True abnormal return per day (spread evenly)
true_ar_daily = true_car / L_event
for i in range(N):
# Generate estimation period residuals
est_residuals = np.random.normal(0, sigma, T_est)
# Generate event window abnormal returns
event_sigma = sigma * np.sqrt(event_variance_multiplier)
event_ar = np.random.normal(true_ar_daily, event_sigma, L_event)
# Create mock model
model = MarketModelResults(
alpha=0.0,
beta=1.0,
sigma=np.std(est_residuals, ddof=2),
r_squared=0.3,
n_obs=T_est,
market_mean=0.0005,
market_var=T_est * 0.0001,
residuals=est_residuals
)
# Create event data
event_data = pd.DataFrame({
'event_time': range(-L_event//2, L_event//2 + L_event%2),
'stock_ret': event_ar + 0.0005, # Add expected return
'market_ret': np.random.normal(0.0005, 0.01, L_event),
'expected_ret': np.full(L_event, 0.0005),
'AR': event_ar
})
results.append({
'model': model,
'event_data': event_data
})
return results
def run_simulation_tests(sim_results: List[Dict], tau1: int, tau2: int) -> Dict:
"""
Run all tests on simulated data.
"""
# Convert to format expected by test functions
class MockResult:
def __init__(self, d):
self.model = d['model']
self.event_data = d['event_data']
mock_results = [MockResult(d) for d in sim_results]
cs = cross_sectional_t_test(mock_results, tau1, tau2)
patell = patell_test(mock_results, tau1, tau2)
bmp = bmp_test(mock_results, tau1, tau2)
return {
'CS_p': cs['p_value'],
'Patell_p': patell['p_value'],
'BMP_p': bmp['p_value']
}
def monte_carlo_power(N_events: int, N_sims: int, true_car: float,
event_var_mult: float = 1.0,
alpha: float = 0.05) -> Dict:
"""
Monte Carlo simulation for test power/size.
"""
rejections = {'CS': 0, 'Patell': 0, 'BMP': 0}
for _ in range(N_sims):
sim_data = simulate_event_study(
N=N_events, T_est=120, L_event=3,
true_car=true_car, sigma=0.02,
event_variance_multiplier=event_var_mult
)
results = run_simulation_tests(sim_data, -1, 1)
if results['CS_p'] < alpha:
rejections['CS'] += 1
if results['Patell_p'] < alpha:
rejections['Patell'] += 1
if results['BMP_p'] < alpha:
rejections['BMP'] += 1
return {
'CS_rejection_rate': rejections['CS'] / N_sims,
'Patell_rejection_rate': rejections['Patell'] / N_sims,
'BMP_rejection_rate': rejections['BMP'] / N_sims
}
print("Monte Carlo Simulation: Test Size and Power")
print("="*70)
print("(This may take a moment...)\n")
N_SIMS = 500 # Reduced for speed; use 1000+ for research
N_EVENTS = 30
# Test size under H0 (true_car = 0)
print("1. SIZE (H0: CAR = 0, No event-induced variance)")
size_results = monte_carlo_power(N_EVENTS, N_SIMS, true_car=0.0, event_var_mult=1.0)
print(f" CS t-test: {size_results['CS_rejection_rate']*100:.1f}% (nominal: 5%)")
print(f" Patell test: {size_results['Patell_rejection_rate']*100:.1f}%")
print(f" BMP test: {size_results['BMP_rejection_rate']*100:.1f}%")
# Test size with event-induced variance
print("\n2. SIZE WITH EVENT-INDUCED VARIANCE (H0: CAR = 0, Variance x2)")
size_eiv = monte_carlo_power(N_EVENTS, N_SIMS, true_car=0.0, event_var_mult=2.0)
print(f" CS t-test: {size_eiv['CS_rejection_rate']*100:.1f}%")
print(f" Patell test: {size_eiv['Patell_rejection_rate']*100:.1f}% <- Over-rejects!")
print(f" BMP test: {size_eiv['BMP_rejection_rate']*100:.1f}% <- Robust")
# Test power
print("\n3. POWER (H1: CAR = 1%)")
power_results = monte_carlo_power(N_EVENTS, N_SIMS, true_car=0.01, event_var_mult=1.0)
print(f" CS t-test: {power_results['CS_rejection_rate']*100:.1f}%")
print(f" Patell test: {power_results['Patell_rejection_rate']*100:.1f}%")
print(f" BMP test: {power_results['BMP_rejection_rate']*100:.1f}%")Monte Carlo Simulation: Test Size and Power
======================================================================
(This may take a moment...)
1. SIZE (H0: CAR = 0, No event-induced variance)
---------------------------------------------------------------------------
ValueError Traceback (most recent call last)
Cell In[31], line 132
130 # Test size under H0 (true_car = 0)
131 print("1. SIZE (H0: CAR = 0, No event-induced variance)")
--> 132 size_results = monte_carlo_power(N_EVENTS, N_SIMS, true_car=0.0, event_var_mult=1.0)
133 print(f" CS t-test: {size_results['CS_rejection_rate']*100:.1f}% (nominal: 5%)")
134 print(f" Patell test: {size_results['Patell_rejection_rate']*100:.1f}%")
Cell In[31], line 101, in monte_carlo_power(N_events, N_sims, true_car, event_var_mult, alpha)
98 rejections = {'CS': 0, 'Patell': 0, 'BMP': 0}
100 for _ in range(N_sims):
--> 101 sim_data = simulate_event_study(
102 N=N_events, T_est=120, L_event=3,
103 true_car=true_car, sigma=0.02,
104 event_variance_multiplier=event_var_mult
105 )
107 results = run_simulation_tests(sim_data, -1, 1)
109 if results['CS_p'] < alpha:
Cell In[31], line 53, in simulate_event_study(N, T_est, L_event, true_car, sigma, event_variance_multiplier)
41 model = MarketModelResults(
42 alpha=0.0,
43 beta=1.0,
(...)
49 residuals=est_residuals
50 )
52 # Create event data
---> 53 event_data = pd.DataFrame({
54 'event_time': range(-L_event//2, L_event//2 + L_event%2),
55 'stock_ret': event_ar + 0.0005, # Add expected return
56 'market_ret': np.random.normal(0.0005, 0.01, L_event),
57 'expected_ret': np.full(L_event, 0.0005),
58 'AR': event_ar
59 })
61 results.append({
62 'model': model,
63 'event_data': event_data
64 })
66 return results
File ~\AppData\Local\anaconda3\Lib\site-packages\pandas\core\frame.py:778, in DataFrame.__init__(self, data, index, columns, dtype, copy)
772 mgr = self._init_mgr(
773 data, axes={"index": index, "columns": columns}, dtype=dtype, copy=copy
774 )
776 elif isinstance(data, dict):
777 # GH#38939 de facto copy defaults to False only in non-dict cases
--> 778 mgr = dict_to_mgr(data, index, columns, dtype=dtype, copy=copy, typ=manager)
779 elif isinstance(data, ma.MaskedArray):
780 from numpy.ma import mrecords
File ~\AppData\Local\anaconda3\Lib\site-packages\pandas\core\internals\construction.py:503, in dict_to_mgr(data, index, columns, dtype, typ, copy)
499 else:
500 # dtype check to exclude e.g. range objects, scalars
501 arrays = [x.copy() if hasattr(x, "dtype") else x for x in arrays]
--> 503 return arrays_to_mgr(arrays, columns, index, dtype=dtype, typ=typ, consolidate=copy)
File ~\AppData\Local\anaconda3\Lib\site-packages\pandas\core\internals\construction.py:114, in arrays_to_mgr(arrays, columns, index, dtype, verify_integrity, typ, consolidate)
111 if verify_integrity:
112 # figure out the index, if necessary
113 if index is None:
--> 114 index = _extract_index(arrays)
115 else:
116 index = ensure_index(index)
File ~\AppData\Local\anaconda3\Lib\site-packages\pandas\core\internals\construction.py:677, in _extract_index(data)
675 lengths = list(set(raw_lengths))
676 if len(lengths) > 1:
--> 677 raise ValueError("All arrays must be of the same length")
679 if have_dicts:
680 raise ValueError(
681 "Mixing dicts with non-Series may lead to ambiguous ordering."
682 )
ValueError: All arrays must be of the same lengthSource
# Power curve as function of true CAR
print("\nGenerating power curves (this may take a minute)...")
true_cars = [0, 0.005, 0.01, 0.015, 0.02, 0.025, 0.03]
power_data = {'CAR': [], 'CS': [], 'Patell': [], 'BMP': []}
for car in true_cars:
result = monte_carlo_power(N_EVENTS, 300, true_car=car)
power_data['CAR'].append(car * 100)
power_data['CS'].append(result['CS_rejection_rate'] * 100)
power_data['Patell'].append(result['Patell_rejection_rate'] * 100)
power_data['BMP'].append(result['BMP_rejection_rate'] * 100)
print(f" CAR = {car*100:.1f}%: CS={result['CS_rejection_rate']*100:.0f}%, "
f"Patell={result['Patell_rejection_rate']*100:.0f}%, "
f"BMP={result['BMP_rejection_rate']*100:.0f}%")
# Plot power curves
fig, ax = plt.subplots(figsize=(10, 6))
ax.plot(power_data['CAR'], power_data['CS'], 'b-o', label='CS t-test', markersize=8)
ax.plot(power_data['CAR'], power_data['Patell'], 'g-s', label='Patell', markersize=8)
ax.plot(power_data['CAR'], power_data['BMP'], 'r-^', label='BMP', markersize=8)
ax.axhline(5, color='gray', linestyle='--', alpha=0.7, label='5% significance')
ax.axhline(80, color='gray', linestyle=':', alpha=0.7, label='80% power target')
ax.set_xlabel('True CAR (%)', fontsize=12)
ax.set_ylabel('Rejection Rate (%)', fontsize=12)
ax.set_title(f'Power Curves (N={N_EVENTS} events, 5% significance)', fontsize=14)
ax.legend()
ax.grid(True, alpha=0.3)
ax.set_ylim(0, 100)
plt.tight_layout()
plt.show()
Generating power curves (this may take a minute)...
---------------------------------------------------------------------------
ValueError Traceback (most recent call last)
Cell In[33], line 8
5 power_data = {'CAR': [], 'CS': [], 'Patell': [], 'BMP': []}
7 for car in true_cars:
----> 8 result = monte_carlo_power(N_EVENTS, 300, true_car=car)
9 power_data['CAR'].append(car * 100)
10 power_data['CS'].append(result['CS_rejection_rate'] * 100)
Cell In[31], line 101, in monte_carlo_power(N_events, N_sims, true_car, event_var_mult, alpha)
98 rejections = {'CS': 0, 'Patell': 0, 'BMP': 0}
100 for _ in range(N_sims):
--> 101 sim_data = simulate_event_study(
102 N=N_events, T_est=120, L_event=3,
103 true_car=true_car, sigma=0.02,
104 event_variance_multiplier=event_var_mult
105 )
107 results = run_simulation_tests(sim_data, -1, 1)
109 if results['CS_p'] < alpha:
Cell In[31], line 53, in simulate_event_study(N, T_est, L_event, true_car, sigma, event_variance_multiplier)
41 model = MarketModelResults(
42 alpha=0.0,
43 beta=1.0,
(...)
49 residuals=est_residuals
50 )
52 # Create event data
---> 53 event_data = pd.DataFrame({
54 'event_time': range(-L_event//2, L_event//2 + L_event%2),
55 'stock_ret': event_ar + 0.0005, # Add expected return
56 'market_ret': np.random.normal(0.0005, 0.01, L_event),
57 'expected_ret': np.full(L_event, 0.0005),
58 'AR': event_ar
59 })
61 results.append({
62 'model': model,
63 'event_data': event_data
64 })
66 return results
File ~\AppData\Local\anaconda3\Lib\site-packages\pandas\core\frame.py:778, in DataFrame.__init__(self, data, index, columns, dtype, copy)
772 mgr = self._init_mgr(
773 data, axes={"index": index, "columns": columns}, dtype=dtype, copy=copy
774 )
776 elif isinstance(data, dict):
777 # GH#38939 de facto copy defaults to False only in non-dict cases
--> 778 mgr = dict_to_mgr(data, index, columns, dtype=dtype, copy=copy, typ=manager)
779 elif isinstance(data, ma.MaskedArray):
780 from numpy.ma import mrecords
File ~\AppData\Local\anaconda3\Lib\site-packages\pandas\core\internals\construction.py:503, in dict_to_mgr(data, index, columns, dtype, typ, copy)
499 else:
500 # dtype check to exclude e.g. range objects, scalars
501 arrays = [x.copy() if hasattr(x, "dtype") else x for x in arrays]
--> 503 return arrays_to_mgr(arrays, columns, index, dtype=dtype, typ=typ, consolidate=copy)
File ~\AppData\Local\anaconda3\Lib\site-packages\pandas\core\internals\construction.py:114, in arrays_to_mgr(arrays, columns, index, dtype, verify_integrity, typ, consolidate)
111 if verify_integrity:
112 # figure out the index, if necessary
113 if index is None:
--> 114 index = _extract_index(arrays)
115 else:
116 index = ensure_index(index)
File ~\AppData\Local\anaconda3\Lib\site-packages\pandas\core\internals\construction.py:677, in _extract_index(data)
675 lengths = list(set(raw_lengths))
676 if len(lengths) > 1:
--> 677 raise ValueError("All arrays must be of the same length")
679 if have_dicts:
680 raise ValueError(
681 "Mixing dicts with non-Series may lead to ambiguous ordering."
682 )
ValueError: All arrays must be of the same length10. Practical Guidelines¶
Which Test to Use?¶
| Situation | Recommended Test | Reason |
|---|---|---|
| General use | BMP | Robust to event-induced variance |
| No event clustering | Patell | More powerful when assumptions hold |
| Event clustering | Kolari-Pynnönen | Adjusts for cross-correlation |
| Very small sample (N<10) | Cross-sectional t | Simple, interpretable |
| Heterogeneous firms | BMP or KP | Account for variance differences |
Reporting Best Practices¶
Report multiple tests for robustness
Include both CAAR and test statistics
Note sample size and any clustering
Report results for multiple windows
Discuss economic vs. statistical significance
Source
def create_publication_table(event_results: List[EventStudyResult],
windows: List[Tuple[int, int]]) -> pd.DataFrame:
"""
Create a publication-ready results table.
"""
rows = []
for tau1, tau2 in windows:
cars = [calculate_car(r.event_data, tau1, tau2) for r in event_results]
caar = np.mean(cars)
cs = cross_sectional_t_test(event_results, tau1, tau2)
patell = patell_test(event_results, tau1, tau2)
bmp = bmp_test(event_results, tau1, tau2)
def sig(p):
if p < 0.01: return '***'
elif p < 0.05: return '**'
elif p < 0.10: return '*'
return ''
rows.append({
'Window': f'[{tau1:+d}, {tau2:+d}]',
'N': len(event_results),
'CAAR (%)': f"{caar*100:+.2f}",
'CS t-stat': f"{cs['t_stat']:.2f}{sig(cs['p_value'])}",
'Patell Z': f"{patell['Z_stat']:.2f}{sig(patell['p_value'])}",
'BMP t-stat': f"{bmp['t_stat']:.2f}{sig(bmp['p_value'])}"
})
return pd.DataFrame(rows)
# Generate publication table
print("\n" + "="*90)
print("PUBLICATION-READY RESULTS TABLE")
print("="*90)
print("\nTable: Cumulative Average Abnormal Returns Around Earnings Announcements")
print("Sample: Tech companies, Q2-Q3 2023 earnings")
print("-"*90)
pub_table = create_publication_table(event_results, windows)
print(pub_table.to_string(index=False))
print("-"*90)
print("Notes: ***, **, * indicate significance at 1%, 5%, 10% levels.")
print("CS = cross-sectional t-test; Patell = Patell (1976) standardized test;")
print("BMP = Boehmer et al. (1991) standardized cross-sectional test.")
==========================================================================================
PUBLICATION-READY RESULTS TABLE
==========================================================================================
Table: Cumulative Average Abnormal Returns Around Earnings Announcements
Sample: Tech companies, Q2-Q3 2023 earnings
------------------------------------------------------------------------------------------
Window N CAAR (%) CS t-stat Patell Z BMP t-stat
[-1, +1] 10 +0.96 0.48 0.55 0.27
[+0, +0] 10 +0.60 1.55 0.79 1.41
[-5, +5] 10 -0.76 -0.37 -0.76 -0.58
[-10, +10] 10 -5.23 -2.08* -1.86* -1.79
------------------------------------------------------------------------------------------
Notes: ***, **, * indicate significance at 1%, 5%, 10% levels.
CS = cross-sectional t-test; Patell = Patell (1976) standardized test;
BMP = Boehmer et al. (1991) standardized cross-sectional test.
11. Exercises¶
Exercise 1: Sample Size Sensitivity¶
Using Monte Carlo simulation, examine how test power changes with sample size (N = 10, 30, 50, 100).
Exercise 2: Window Length Effects¶
How does test power change as you lengthen the event window? Why?
Exercise 3: Skewness Adjustment¶
Implement a skewness-adjusted t-test that accounts for non-normality in CARs.
Exercise 4: Calendar-Time Portfolio¶
Implement a calendar-time portfolio approach that forms portfolios of event firms and regresses on market factors.
Source
# Exercise 3 Template: Skewness-Adjusted t-Test
def skewness_adjusted_t_test(event_results: List[EventStudyResult],
tau1: int, tau2: int) -> Dict:
"""
Hall (1992) skewness-adjusted t-test.
Adjusts for skewness in the CAR distribution.
"""
cars = np.array([calculate_car(r.event_data, tau1, tau2) for r in event_results])
N = len(cars)
# Standard t-test components
mean_car = np.mean(cars)
std_car = np.std(cars, ddof=1)
se = std_car / np.sqrt(N)
t_standard = mean_car / se if se > 0 else 0
# Skewness
skew = stats.skew(cars)
# Hall (1992) adjustment
# t_adj = t + (1/6) * skew * (t^2 + 2) / sqrt(N)
t_adjusted = t_standard + (1/6) * skew * (t_standard**2 + 2) / np.sqrt(N)
p_value = 2 * (1 - stats.t.cdf(abs(t_adjusted), df=N-1))
return {
'test_name': 'Skewness-Adjusted t',
'N': N,
'CAAR': mean_car,
'skewness': skew,
't_standard': t_standard,
't_adjusted': t_adjusted,
'p_value': p_value
}
# Test the skewness-adjusted t-test
print("Skewness-Adjusted t-Test:")
print("="*50)
for tau1, tau2 in [(-1, +1), (0, 0)]:
result = skewness_adjusted_t_test(event_results, tau1, tau2)
print(f"\nWindow [{tau1:+d}, {tau2:+d}]:")
print(f" Skewness = {result['skewness']:.3f}")
print(f" Standard t = {result['t_standard']:.3f}")
print(f" Adjusted t = {result['t_adjusted']:.3f}")
print(f" p-value = {result['p_value']:.4f}")Skewness-Adjusted t-Test:
==================================================
Window [-1, +1]:
Skewness = -0.507
Standard t = 0.475
Adjusted t = 0.416
p-value = 0.6873
Window [+0, +0]:
Skewness = -0.001
Standard t = 1.550
Adjusted t = 1.550
p-value = 0.1556
12. Summary¶
In this session, we covered:
Cross-Sectional t-Test: Simple but assumes homogeneous variance
Time-Series t-Test: Uses estimation period variance
Patell (1976) Test: Standardizes ARs, more powerful but assumes no event-induced variance
BMP (1991) Test: Robust to event-induced variance changes
Kolari-Pynnönen (2010): Adjusts for cross-correlation
Monte Carlo Analysis: Understanding test size and power
Key Takeaways¶
BMP is the recommended default for most applications
Event-induced variance is common and Patell over-rejects
Cross-correlation from event clustering requires adjustment
Report multiple tests for robustness
Consider economic significance alongside statistical significance
Coming Up Next¶
Session 5: Statistical Inference II - Non-Parametric and Robust Tests will cover:
Sign test and generalized sign test
Rank tests (Corrado, Corrado-Zivney)
Bootstrap methods
Dealing with non-normality
13. References¶
Core Methodology¶
Patell, J. M. (1976). Corporate forecasts of earnings per share and stock price behavior. Journal of Accounting Research, 14(2), 246-276.
Boehmer, E., Musumeci, J., & Poulsen, A. B. (1991). Event-study methodology under conditions of event-induced variance. Journal of Financial Economics, 30(2), 253-272.
Kolari, J. W., & Pynnönen, S. (2010). Event study testing with cross-sectional correlation of abnormal returns. Review of Financial Studies, 23(11), 3996-4025.
Power and Specification¶
Brown, S. J., & Warner, J. B. (1985). Using daily stock returns: The case of event studies. Journal of Financial Economics, 14(1), 3-31.
Campbell, J. Y., Lo, A. W., & MacKinlay, A. C. (1997). The Econometrics of Financial Markets. Princeton University Press.